The candidate plasticity gene cpg2 transcript is a product of the Syne1 gene, a large gene encoding multiple proteins with different functions. The cpg2 expression pattern, activity-dependent regulation, and sub-cellular localization of its protein product are all consistent with a role in synaptic plasticity (Cottrell et al., Neuron 2004). The CPG2 protein is localized to an endocytic zone adjacent to excitatory synapses in dendritic spines. At this site, CPG2 plays a role in constitutive as well as activity-dependent endocytosis of synaptic glutamate receptors (Cottrell et al., Neuron 2004). At the synapse, clathrin mediated endocytosis (CME) of AMPA-type glutamate receptors (AMPARs) from the postsynaptic membrane is thought to be the substrate for various forms of plasticity, including long-term depression (LTD).
The actin cytoskeleton is critical for CME, yet the mechanistic understanding of its interaction with the endocytic process has been elusive. We have identified CPG2 as an F-actin binding partner that functionally mediates interaction of the spine cytoskeleton with postsynaptic endocytosis. Further, the regulation of CPG2/F-actin association by PKA provides a gateway for cellular control of synaptic receptor internalization through second messenger signaling pathways (Loebrich et al. PNAS 2013). Recent identification of human SYNE1 as a risk locus for bipolar disorder suggests that CPG2 could play a role in synaptic dysfunction underlying neuropsychiatric disease. We are currently identifying human homologues of cpg2 and testing if naturally occurring mutations in these transcripts alter glutamatergic synaptic function. In parallel, we are working to identify functional modules in the CPG2 protein that mediate distinct protein interactions associated with glutamate receptor endocytosis.
CPG2 and Bipolar Disorder
Bipolar Disorder (BD) is a debilitating and heritable mood disorder characterized by recurrent episodes of mania and depression. Drug treatment is often unsuccessful, and full recovery between episodes is not achieved in all patients. As a consequence, BD is a leading cause of disability, loss of life through high suicide rates and contributes to the emotional and financial burden for families and society. There have been few consistent leads in terms of the biological underpinnings of BD. Growing evidence suggests that the glutamatergic system is central to the neurobiology of mood disorder. Postmortem studies have shown decreased NMDA glutamate receptor (NMDAR) expression in cortical and hippocampal tissue from BD patients. Also, decreased expression of AMPA type glutamate receptors (AMPAR) has been found in cortical and striatal BD brain tissue. Another tantalizing link to glutamatergic synaptic function is through the effects of pharmacological treatments of mood disorders. Lithium and valproate, the mainline treatments for BD, affect AMPAR activation and trafficking, and single-nucleotide polymorphisms (SNPs) in GRIA2, which encodes the GluA2 subunit of AMPARs, have been associated with time to recurrence of mood episodes in BD patients on lithium.
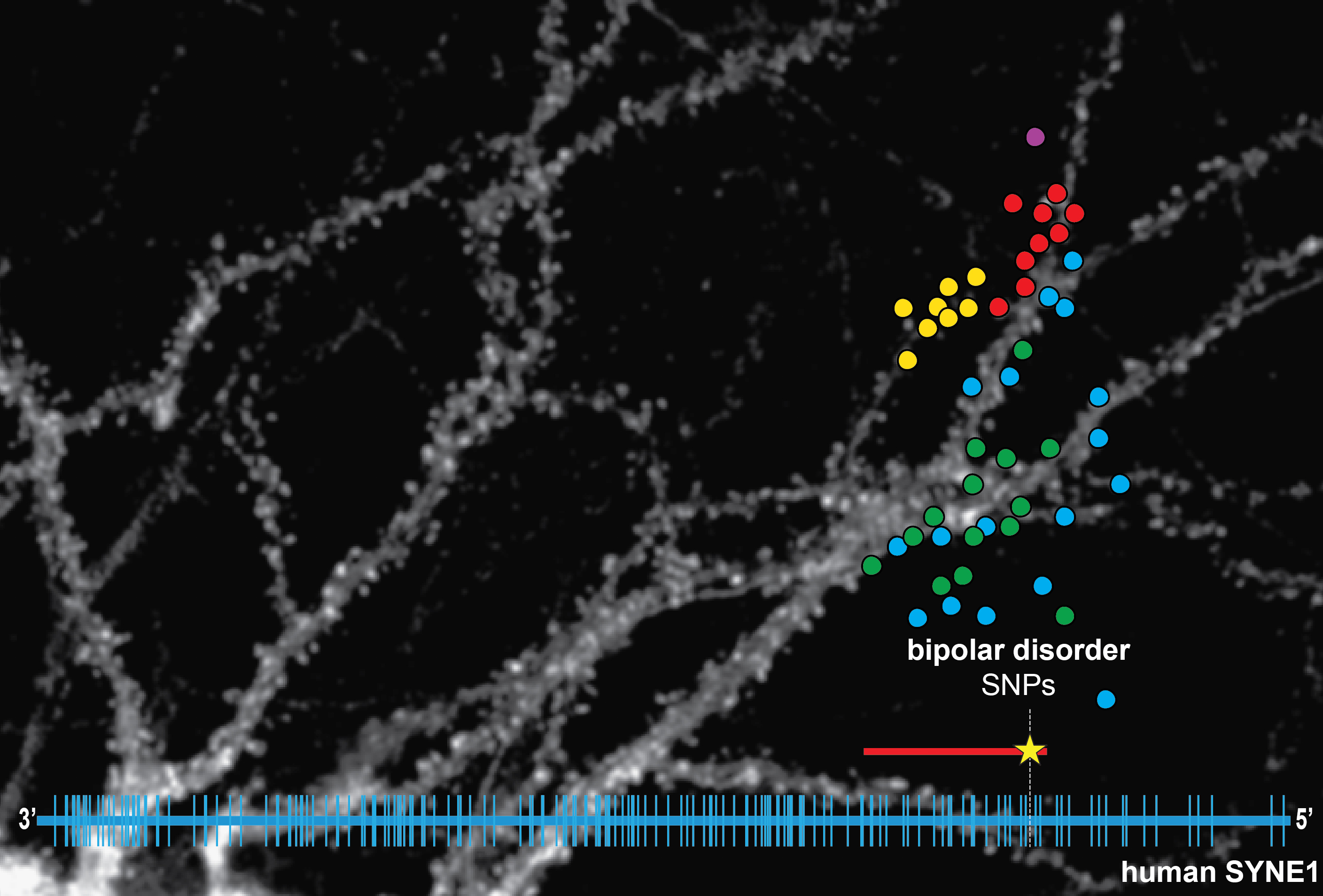
Recent human genome-wide association studies (GWAS) for genes that confer risk for schizophrenia, BD and major depression have revealed an association between SNPs in the SYNE1 gene and increased risk of BD. The BD association signal spans, almost exclusively, parts of human SYNE1 that encode CPG2. Given the critical role that CPG2 plays in glutamate receptor internalization our working hypothesis is that genetic variations in CPG2 affect its interaction with cytoskeletal/endocytic binding partners, thus influencing the efficacy of glutamatergic transmission. Through collaboration with the Stanley Center for Psychiatric Research (at the Broad Institute of MIT & Harvard in Cambridge, MA), we screened deep-sequencing data of the SYNE1 gene in patient cohorts and identified additional SNPs that are potentially disease related in the CPG2 locus of human SYNE1. By transcript mapping of this locus we showed the existence of Cpg2 transcripts and their protein products in human brain. We found a strong conservation of CPG2 function between rat and human, opening the door to testing the effect of missense SNPs identified by BD patient exome sequencing on neuronal function. We have a unique opportunity to link BD associated variation with an underlying cellular dysfunction, an opportunity afforded by very few GWAS hits in human genetics for neuropsychiatric disorders to date.
CPG2 Functional Pathways
The number of glutamate receptors at the post-synaptic membrane determines the strength of excitatory neurotransmission by altering the sensitivity of the post-synaptic cell to neurotransmitter release. We are investigating the molecular mechanisms regulating glutamate receptor surface expression in order to understand excitatory synaptic plasticity. Candidate Plasticity Gene 2 (CPG2) encodes a protein that localizes to the post-synaptic spines of excitatory neurons where it regulates the internalization of glutamate receptors. CPG2 has a modular structure with multiple protein interaction domains suggesting it functions as a large adaptor complex that can mediate the interaction of varied synaptic components involved in regulation of glutamate receptor cycling at the synapse.
We have previously shown that CPG2 facilitates receptor endocytosis through interactions with f-actin and this interaction is mediated by phosphorylation of CPG2 by protein kinase A, suggesting a mechanism for second messenger regulation of glutamate receptor internalization (Loebrich et al., PNAS 2013). The goal of my project is to identify protein-protein interactions between CPG2 and other potential binding partners or post-translational modifiers that may function in this pathway, in particular ones that interface with the endocytic machinery. By studying these processes, we hope to reveal important cellular mechanisms underlying synaptic circuitry modulation and brain plasticity in response to fluctuations in neuronal activity.